Foundation models for generalist medical artificial intelligence
Abstract
The exceptionally rapid development of highly flexible, reusable artificial intelligence (AI) models is likely to usher in newfound capabilities in medicine. We propose a new paradigm for medical AI, which we refer to as generalist medical AI (GMAI). GMAI models will be capable of carrying out a diverse set of tasks using very little or no task-specific labelled data. Built through self-supervision on large, diverse datasets, GMAI will flexibly interpret different combinations of medical modalities, including data from imaging, electronic health records, laboratory results, genomics, graphs or medical text. Models will in turn produce expressive outputs such as free-text explanations, spoken recommendations or image annotations that demonstrate advanced medical reasoning abilities. Here we identify a set of high-impact potential applications for GMAI and lay out specific technical capabilities and training datasets necessary to enable them. We expect that GMAI-enabled applications will challenge current strategies for regulating and validating AI devices for medicine and will shift practices associated with the collection of large medical datasets.
Main
Foundation models—the latest generation of AI models—are trained on massive, diverse datasets and can be applied to numerous downstream tasks1. Individual models can now achieve state-of-the-art performance on a wide variety of problems, ranging from answering questions about texts to describing images and playing video games2,3,4. This versatility represents a stark change from the previous generation of AI models, which were designed to solve specific tasks, one at a time.
Driven by growing datasets, increases in model size and advances in model architectures, foundation models offer previously unseen abilities. For example, in 2020 the language model GPT-3 unlocked a new capability: in-context learning, through which the model carried out entirely new tasks that it had never explicitly been trained for, simply by learning from text explanations (or ‘prompts’) containing a few examples5. Additionally, many recent foundation models are able to take in and output combinations of different data modalities4,6. For example, the recent Gato model can chat, caption images, play video games and control a robot arm and has thus been described as a generalist agent2. As certain capabilities emerge only in the largest models, it remains challenging to predict what even larger models will be able to accomplish7.
Although there have been early efforts to develop medical foundation models8,9,10,11, this shift has not yet widely permeated medical AI, owing to the difficulty of accessing large, diverse medical datasets, the complexity of the medical domain and the recency of this development. Instead, medical AI models are largely still developed with a task-specific approach to model development. For instance, a chest X-ray interpretation model may be trained on a dataset in which every image has been explicitly labelled as positive or negative for pneumonia, probably requiring substantial annotation effort. This model would only detect pneumonia and would not be able to carry out the complete diagnostic exercise of writing a comprehensive radiology report. This narrow, task-specific approach produces inflexible models, limited to carrying out tasks predefined by the training dataset and its labels. In current practice, such models typically cannot adapt to other tasks (or even to different data distributions for the same task) without being retrained on another dataset. Of the more than 500 AI models for clinical medicine that have received approval by the Food and Drug Administration, most have been approved for only 1 or 2 narrow tasks12.
Here we outline how recent advances in foundation model research can disrupt this task-specific paradigm. These include the rise of multimodal architectures13 and self-supervised learning techniques14 that dispense with explicit labels (for example, language modelling15 and contrastive learning16), as well as the advent of in-context learning capabilities5.
These advances will instead enable the development of GMAI, a class of advanced medical foundation models. ‘Generalist’ implies that they will be widely used across medical applications, largely replacing task-specific models.
Inspired directly by foundation models outside medicine, we identify three key capabilities that distinguish GMAI models from conventional medical AI models (Fig. 1). First, adapting a GMAI model to a new task will be as easy as describing the task in plain English (or another language). Models will be able to solve previously unseen problems simply by having new tasks explained to them (dynamic task specification), without needing to be retrained3,5. Second, GMAI models can accept inputs and produce outputs using varying combinations of data modalities (for example, can take in images, text, laboratory results or any combination thereof). This flexible interactivity contrasts with the constraints of more rigid multimodal models, which always use predefined sets of modalities as input and output (for example, must always take in images, text and laboratory results together). Third, GMAI models will formally represent medical knowledge, allowing them to reason through previously unseen tasks and use medically accurate language to explain their outputs.
We list concrete strategies for achieving this paradigm shift in medical AI. Furthermore, we describe a set of potentially high-impact applications that this new generation of models will enable. Finally, we point out core challenges that must be overcome for GMAI to deliver the clinical value it promises.
The potential of generalist models in medical AI
GMAI models promise to solve more diverse and challenging tasks than current medical AI models, even while requiring little to no labels for specific tasks. Of the three defining capabilities of GMAI, two enable flexible interactions between the GMAI model and the user: first, the ability to carry out tasks that are dynamically specified; and second, the ability to support flexible combinations of data modalities. The third capability requires that GMAI models formally represent medical domain knowledge and leverage it to carry out advanced medical reasoning. Recent foundation models already exhibit individual aspects of GMAI, by flexibly combining several modalities2 or making it possible to dynamically specify a new task at test time5, but substantial advances are still required to build a GMAI model with all three capabilities. For example, existing models that show medical reasoning abilities (such as GPT-3 or PaLM) are not multimodal and do not yet generate reliably factual statements.
Flexible interactions
GMAI offers users the ability to interact with models through custom queries, making AI insights easier for different audiences to understand and offering unprecedented flexibility across tasks and settings. In current practice, AI models typically handle a narrow set of tasks and produce a rigid, predetermined set of outputs. For example, a current model might detect a specific disease, taking in one kind of image and always outputting the likelihood of that disease. By contrast, a custom query allows users to come up with questions on the fly: “Explain the mass appearing on this head MRI scan. Is it more likely a tumour or an abscess?”. Furthermore, queries can allow users to customize the format of their outputs: “This is a follow-up MRI scan of a patient with glioblastoma. Outline any tumours in red”.
Custom queries will enable two key capabilities—dynamic task specification and multimodal inputs and outputs—as follows.
Dynamic task specification
Custom queries can teach AI models to solve new problems on the fly, dynamically specifying new tasks without requiring models to be retrained. For example, GMAI can answer highly specific, previously unseen questions: “Given this ultrasound, how thick is the gallbladder wall in millimetres?”. Unsurprisingly, a GMAI model may struggle to complete new tasks that involve unknown concepts or pathologies. In-context learning then allows users to teach the GMAI about a new concept with few examples: “Here are the medical histories of ten previous patients with an emerging disease, an infection with the Langya henipavirus. How likely is it that our current patient is also infected with Langya henipavirus?”17.
Multimodal inputs and outputs
Custom queries can allow users to include complex medical information in their questions, freely mixing modalities. For example, a clinician might include multiple images and laboratory results in their query when asking for a diagnosis. GMAI models can also flexibly incorporate different modalities into responses, such as when a user asks for both a text answer and an accompanying visualization. Following previous models such as Gato, GMAI models can combine modalities by turning each modality’s data into ‘tokens’, each representing a small unit (for example, a word in a sentence or a patch in an image) that can be combined across modalities. This blended stream of tokens can then be fed into a transformer architecture18, allowing GMAI models to integrate a given patient’s entire history, including reports, waveform signals, laboratory results, genomic profiles and imaging studies.
Medical domain knowledge
In stark contrast to a clinician, conventional medical AI models typically lack prior knowledge of the medical domain before they are trained for their particular tasks. Instead, they have to rely solely on statistical associations between features of the input data and the prediction target, without having contextual information (for example, about pathophysiological processes). This lack of background makes it harder to train models for specific medical tasks, particularly when data for the tasks are scarce.
GMAI models can address these shortcomings by formally representing medical knowledge. For example, structures such as knowledge graphs can allow models to reason about medical concepts and relationships between them. Furthermore, building on recent retrieval-based approaches, GMAI can retrieve relevant context from existing databases, in the form of articles, images or entire previous cases19,20.
The resulting models can raise self-explanatory warnings: “This patient is likely to develop acute respiratory distress syndrome, because the patient was recently admitted with a severe thoracic trauma and because the patient’s partial pressure of oxygen in the arterial blood has steadily decreased, despite an increased inspired fraction of oxygen”.
As a GMAI model may even be asked to provide treatment recommendations, despite mostly being trained on observational data, the model’s ability to infer and leverage causal relationships between medical concepts and clinical findings will play a key role for clinical applicability21.
Finally, by accessing rich molecular and clinical knowledge, a GMAI model can solve tasks with limited data by drawing on knowledge of related problems, as exemplified by initial works on AI-based drug repurposing22.
Use cases of GMAI
We present six potential use cases for GMAI that target different user bases and disciplines, although our list is hardly exhaustive. Although there have already been AI efforts in these areas, we expect GMAI will enable comprehensive solutions for each problem.
Grounded radiology reports
GMAI enables a new generation of versatile digital radiology assistants, supporting radiologists throughout their workflow and markedly reducing workloads. GMAI models can automatically draft radiology reports that describe both abnormalities and relevant normal findings, while also taking into account the patient’s history. These models can provide further assistance to clinicians by pairing text reports with interactive visualizations, such as by highlighting the region described by each phrase. Radiologists can also improve their understanding of cases by chatting with GMAI models: “Can you highlight any new multiple sclerosis lesions that were not present in the previous image?”.
A solution needs to accurately interpret various radiology modalities, noticing even subtle abnormalities. Furthermore, it must integrate information from a patient’s history, including sources such as indications, laboratory results and previous images, when describing an image. It also needs to communicate with clinicians using multiple modalities, providing both text answers and dynamically annotated images. To do so, it must be capable of visual grounding, accurately pointing out exactly which part of an image supports any statement. Although this may be achieved through supervised learning on expert-labelled images, explainability methods such as Grad-CAM could enable self-supervised approaches, requiring no labelled data23.
Augmented procedures
We anticipate a surgical GMAI model that can assist surgical teams with procedures: “We cannot find the intestinal rupture. Check whether we missed a view of any intestinal section in the visual feed of the last 15 minutes”. GMAI models may carry out visualization tasks, potentially annotating video streams of a procedure in real time. They may also provide information in spoken form, such as by raising alerts when steps of a procedure are skipped or by reading out relevant literature when surgeons encounter rare anatomical phenomena.

This model can also assist with procedures outside the operating room, such as with endoscopic procedures. A model that captures topographic context and reasons with anatomical knowledge can draw conclusions about previously unseen phenomena. For instance, it could deduce that a large vascular structure appearing in a duodenoscopy may indicate an aortoduodenal fistula (that is, an abnormal connection between aorta and the small intestine), despite never having encountered one before (Fig. 2, right panel). GMAI can solve this task by first detecting the vessel, second identifying the anatomical location, and finally considering the neighbouring structures.
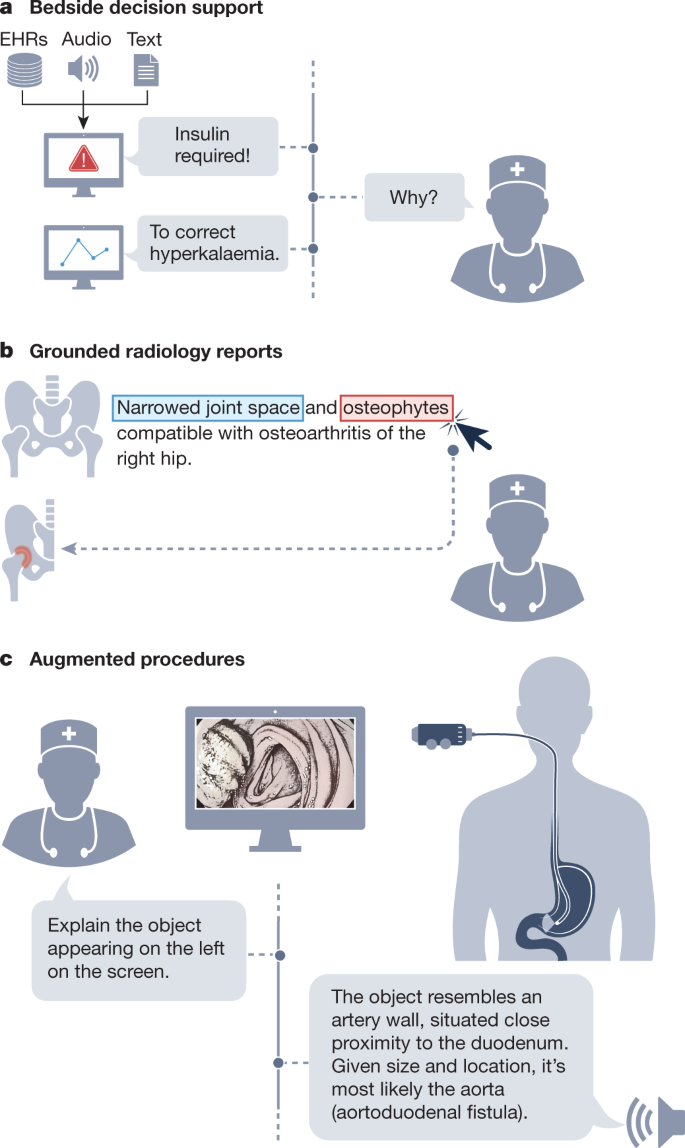
A solution needs to integrate vision, language and audio modalities, using a vision–audio–language model to accept spoken queries and carry out tasks using the visual feed. Vision–language models have already gained traction, and the development of models that incorporate further modalities is merely a question of time24. Approaches may build on previous work that combines language models and knowledge graphs25,26 to reason step-by-step about surgical tasks. Additionally, GMAI deployed in surgical settings will probably face unusual clinical phenomena that cannot be included during model development, owing to their rarity, a challenge known as the long tail of unseen conditions27. Medical reasoning abilities will be crucial for both detecting previously unseen outliers and explaining them, as exemplified in Fig. 2.
Bedside decision support
GMAI enables a new class of bedside clinical decision support tools that expand on existing AI-based early warning systems, providing more detailed explanations as well as recommendations for future care. For example, GMAI models for bedside decision support can leverage clinical knowledge and provide free-text explanations and data summaries: “Warning: This patient is about to go into shock. Her circulation has destabilized in the last 15 minutes <link to data summary>. Recommended next steps: <link to checklist>”.
A solution needs to parse electronic health record (EHR) sources (for example, vital and laboratory parameters, and clinical notes) that involve multiple modalities, including text and numeric time series data. It needs to be able to summarize a patient’s current state from raw data, project potential future states of the patient and recommend treatment decisions. A solution may project how a patient’s condition will change over time, by using language modelling techniques to predict their future textual and numeric records from their previous data. Training datasets may specifically pair EHR time series data with eventual patient outcomes, which can be collected from discharge reports and ICD (International Classification of Diseases) codes. In addition, the model must be able to compare potential treatments and estimate their effects, all while adhering to therapeutic guidelines and other relevant policies. The model can acquire the necessary knowledge through clinical knowledge graphs and text sources such as academic publications, educational textbooks, international guidelines and local policies. Approaches may be inspired by REALM, a language model that answers queries by first retrieving a single relevant document and then extracting the answer from it, making it possible for users to identify the exact source of each answer20.
Interactive note-taking
Documentation represents an integral but labour-intensive part of clinical workflows. By monitoring electronic patient information as well as clinician–patient conversations, GMAI models will preemptively draft documents such as electronic notes and discharge reports for clinicians to merely review, edit and approve. Thus, GMAI can substantially reduce administrative overhead, allowing clinicians to spend more time with patients.
A GMAI solution can draw from recent advances in speech-to-text models28, specializing techniques for medical applications. It must accurately interpret speech signals, understanding medical jargon and abbreviations. Additionally, it must contextualize speech data with information from the EHRs (for example, diagnosis list, vital parameters and previous discharge reports) and then generate free-text notes or reports. It will be essential to obtain consent before recording any interaction with a patient. Even before such recordings are collected in large numbers, early note-taking models may already be developed by leveraging clinician–patient interaction data collected from chat applications.
Chatbots for patients
GMAI has the potential to power new apps for patient support, providing high-quality care even outside clinical settings. For example, GMAI can build a holistic view of a patient’s condition using multiple modalities, ranging from unstructured descriptions of symptoms to continuous glucose monitor readings to patient-provided medication logs. After interpreting these heterogeneous types of data, GMAI models can interact with the patient, providing detailed advice and explanations. Importantly, GMAI enables accessible communication, providing clear, readable or audible information on the patient’s schedule. Whereas similar apps rely on clinicians to offer personalized support at present29, GMAI promises to reduce or even remove the need for human expert intervention, making apps available on a larger scale. As with existing live chat applications, users could still engage with a human counsellor on request.
Building patient-facing chatbots with GMAI raises two special challenges. First, patient-facing models must be able to communicate clearly with non-technical audiences, using simple, clear language without sacrificing the accuracy of the content. Including patient-focused medical texts in training datasets may enable this capability. Second, these models need to work with diverse data collected by patients. Patient-provided data may represent unusual modalities; for example, patients with strict dietary requirements may submit before-and-after photos of their meals so that GMAI models can automatically monitor their food intake. Patient-collected data are also likely to be noisier compared to data from a clinical setting, as patients may be more prone to error or use less reliable devices when collecting data. Again, incorporating relevant data into training can help overcome this challenge. However, GMAI models also need to monitor their own uncertainty and take appropriate action when they do not have enough reliable data.
Text-to-protein generation
GMAI could generate protein amino acid sequences and their three-dimensional structures from textual prompts. Inspired by existing generative models of protein sequences30, such a model could condition its generation on desired functional properties. By contrast, a biomedically knowledgeable GMAI model promises protein design interfaces that are as flexible and easy to use as concurrent text-to-image generative models such as Stable Diffusion or DALL-E31,32. Moreover, by unlocking in-context learning capabilities, a GMAI-based text-to-protein model may be prompted with a handful of example instructions paired with sequences to dynamically define a new generation task, such as the generation of a protein that binds with high affinity to a specified target while fulfilling additional constraints.
There have already been early efforts to develop foundation models for biological sequences33,34, including RFdiffusion, which generates proteins on the basis of simple specifications (for example, a binding target)35. Building on this work, GMAI-based solution can incorporate both language and protein sequence data during training to offer a versatile text interface. A solution could also draw on recent advances in multimodal AI such as CLIP, in which models are jointly trained on paired data of different modalities16. When creating such a training dataset, individual protein sequences must be paired with relevant text passages (for example, from the body of biological literature) that describe the properties of the proteins. Large-scale initiatives, such as UniProt, that map out protein functions for millions of proteins, will be indispensable for this effort36.
Opportunities and challenges of GMAI
GMAI has the potential to affect medical practice by improving care and reducing clinician burnout. Here we detail the overarching advantages of GMAI models. We also describe critical challenges that must be addressed to ensure safe deployment, as GMAI models will operate in particularly high-stakes settings, compared to foundation models in other fields.
Paradigm shifts with GMAI
Controllability
GMAI allows users to finely control the format of its outputs, making complex medical information easier to access and understand. For example, there will be GMAI models that can rephrase natural language responses on request. Similarly, GMAI-provided visualizations may be carefully tailored, such as by changing the viewpoint or labelling important features with text. Models can also potentially adjust the level of domain-specific detail in their outputs or translate them into multiple languages, communicating effectively with diverse users. Finally, GMAI’s flexibility allows it to adapt to particular regions or hospitals, following local customs and policies. Users may need formal instruction on how to query a GMAI model and to use its outputs most effectively.
Adaptability
Existing medical AI models struggle with distribution shifts, in which distributions of data shift owing to changes in technologies, procedures, settings or populations37,38. However, GMAI can keep pace with shifts through in-context learning. For example, a hospital can teach a GMAI model to interpret X-rays from a brand-new scanner simply by providing prompts that show a small set of examples. Thus, GMAI can adapt to new distributions of data on the fly, whereas conventional medical AI models would need to be retrained on an entirely new dataset. At present, in-context learning is observed predominantly in large language models39. To ensure that GMAI can adapt to changes in context, a GMAI model backbone needs to be trained on extremely diverse data from multiple, complementary sources and modalities. For instance, to adapt to emerging variants of coronavirus disease 2019, a successful model can retrieve characteristics of past variants and update them when confronted with new context in a query. For example, a clinician might say, “Check these chest X-rays for Omicron pneumonia. Compared to the Delta variant, consider infiltrates surrounding the bronchi and blood vessels as indicative signs”40.
Although users can manually adjust model behaviour through prompts, there may also be a role for new techniques to automatically incorporate human feedback. For example, users may be able to rate or comment on each output from a GMAI model, much as users rate outputs of ChatGPT (released by OpenAI in 2022), an AI-powered chat interface. Such feedback can then be used to improve model behaviour, following the example of InstructGPT, a model created by using human feedback to refine GPT-3 through reinforcement learning41.
Applicability
Large-scale AI models already serve as the foundation for numerous downstream applications. For instance, within months after its release, GPT-3 powered more than 300 apps across various industries42. As a promising early example of a medical foundation model, CheXzero can be applied to detect dozens of diseases in chest X-rays without being trained on explicit labels for these diseases9. Likewise, the shift towards GMAI will drive the development and release of large-scale medical AI models with broad capabilities, which will form the basis for various downstream clinical applications. Many applications will interface with the GMAI model itself, directly using its final outputs. Others may use intermediate numeric representations, which GMAI models naturally generate in the process of producing outputs, as inputs for small specialist models that can be cheaply built for specific tasks. However, this flexible applicability can act as a double-edged sword, as any failure mode that exists in the foundation model will be propagated widely throughout the downstream applications.
Challenges of GMAI
Validation
GMAI models will be uniquely difficult to validate, owing to their unprecedented versatility. At present, AI models are designed for specific tasks, so they need to be validated only for those predefined use cases (for example, diagnosing a particular type of cancer from a brain MRI). However, GMAI models can carry out previously unseen tasks set forth by an end user for the first time (for example, diagnosing any disease in a brain MRI), so it is categorically more challenging to anticipate all of their failure modes. Developers and regulators will be responsible for explaining how GMAI models have been tested and what use cases they have been approved for. GMAI interfaces themselves should be designed to raise ‘off-label usage’ warnings on entering uncharted territories, instead of confidently fabricating inaccurate information. More generally, GMAI’s uniquely broad capabilities require regulatory foresight, demanding that institutional and governmental policies adapt to the new paradigm, and will also reshape insurance arrangements and liability assignment.
Verification
Compared to conventional AI models, GMAI models can handle unusually complex inputs and outputs, making it more difficult for clinicians to determine their correctness. For example, conventional models may consider only an imaging study or a whole-slide image when classifying a patient’s cancer. In each case, a sole radiologist or pathologist could verify whether the model’s outputs are correct. However, a GMAI model may consider both kinds of inputs and may output an initial classification, a recommendation for treatment and a multimodal justification involving visualizations, statistical analyses and references to the literature. In this case, a multidisciplinary panel (consisting of radiologists, pathologists, oncologists and additional specialists) may be needed to judge the GMAI’s output. Fact-checking GMAI outputs therefore represents a serious challenge, both during validation and after models are deployed.
Creators can make it easier to verify GMAI outputs by incorporating explainability techniques. For example, a GMAI’s outputs might include clickable links to supporting passages in the literature, allowing clinicians to more efficiently verify GMAI predictions. Other strategies for fact-checking a model’s output without human expertise have recently been proposed43. Finally, it is vitally important that GMAI models accurately express uncertainty, thereby preventing overconfident statements in the first place.
Social biases
Previous work has already shown that medical AI models can perpetuate biases and cause harm to marginalized populations. They can acquire biases during training, when datasets either underrepresent certain groups of patients or contain harmful correlations44,45. These risks will probably be even more pronounced when developing GMAI. The unprecedented scale and complexity of the necessary training datasets will make it difficult to ensure that they are free of undesirable biases. Although biases already pose a challenge for conventional AI in health, they are of particular relevance for GMAI as a recent large-scale evaluation showed that social bias can increase with model scale46.
GMAI models must be thoroughly validated to ensure that they do not underperform on particular populations such as minority groups. Furthermore, models will need to undergo continuous auditing and regulation even after deployment, as new issues will arise as models encounter new tasks and settings. Prize-endowed competitions could incentivize the AI community to further scrutinize GMAI models. For instance, participants might be rewarded for finding prompts that produce harmful content or expose other failure modes. Swiftly identifying and fixing biases must be an utmost priority for developers, vendors and regulators.
Privacy
The development and use of GMAI models poses serious risks to patient privacy. GMAI models may have access to a rich set of patient characteristics, including clinical measurements and signals, molecular signatures and demographic information as well as behavioural and sensory tracking data. Furthermore, GMAI models will probably use large architectures, but larger models are more prone to memorizing training data and directly repeating it to users47. As a result, there is a serious risk that GMAI models could expose sensitive patient data in training datasets. By means of deidentification and limiting the amount of information collected for individual patients, the damage caused by exposed data can be reduced.
However, privacy concerns are not limited to training data, as deployed GMAI models may also expose data from current patients. Prompt attacks can trick models such as GPT-3 into ignoring previous instructions48. As an example, imagine that a GMAI model has been instructed never to reveal patient information to uncredentialed users. A malicious user could force the model to ignore that instruction to extract sensitive data.
Scale
Recent foundation models have increased markedly in size, driving up costs associated with data collection and model training. Models of this scale require massive training datasets that, in the case of GPT-3, contain hundreds of billions of tokens and are expensive to collect. Furthermore, PaLM, a 540-billion-parameter model developed by Google, required an estimated 8.4 million hours’ worth of tensor processing unit v4 chips for training, using roughly 3,000 to 6,000 chips at a time, amounting to millions of dollars in computational costs49. Additionally, developing such large models brings a substantial environmental cost, as training each model has been estimated to generate up to hundreds of tons of CO2 equivalent50.
These costs raise the question of how large datasets and models should be. One recent study established a link between dataset size and model size, recommending 20 times more tokens than parameters for optimal performance, yet existing foundation models were successfully trained with a lower token-to-parameter ratio51. It thus remains difficult to estimate how large models and datasets must be when developing GMAI models, especially because the necessary scale depends heavily on the particular medical use case.
Data collection will pose a particular challenge for GMAI development, owing to the need for unprecedented amounts of medical data. Existing foundation models are typically trained on heterogeneous data obtained by crawling the web, and such general-purpose data sources can potentially be used to pretrain GMAI models (that is, carry out an initial preparatory round of training). Although these datasets do not focus on medicine, such pretraining can equip GMAI models with useful capabilities. For example, by drawing on medical texts present within their training datasets, general-purpose models such as Flan-PaLM or ChatGPT can accurately answer medical questions, achieving passing scores on the United States Medical Licensing Exam10,52,53. Nevertheless, GMAI model development will probably also require massive datasets that specifically focus on the medical domain and its modalities. These datasets must be diverse, anonymized and organized in compatible formats, and procedures for collecting and sharing data will need to comply with heterogeneous policies across institutions and regions. Although gathering such large datasets will pose a substantial challenge, these data will generally not require costly expert labels, given the success of self-supervision9,54. Additionally, multimodal self-supervision techniques can be used to train models on multiple datasets containing measurements from a few modalities each, reducing the need for large, expensive datasets that contain measurements from many modalities per patient. In other words, a model can be trained on one dataset with EHR and MRI data and a second with EHR and genomic data, without requiring a large dataset that contains EHR, MRI and genomic data, jointly. Large-scale data-sharing efforts, such as the MIMIC (Medical Information Mart for Intensive Care) database55 or the UK Biobank56, will play a critical role in GMAI, and they should be extended to underrepresented countries to create larger, richer and more inclusive training datasets.
The size of GMAI models will also cause technical challenges. In addition to being costly to train, GMAI models can be challenging to deploy, requiring specialized, high-end hardware that may be difficult for hospitals to access. For certain use cases (for example, chatbots), GMAI models can be stored on central compute clusters maintained by organizations with deep technical expertise, as DALL-E or GPT-3 are. However, other GMAI models may need to be deployed locally in hospitals or other medical settings, removing the need for a stable network connection and keeping sensitive patient data on-site. In these cases, model size may need to be reduced through techniques such as knowledge distillation, in which large-scale models teach smaller models that can be more easily deployed under practical constraints57.
Conclusion
Foundation models have the potential to transform healthcare. The class of advanced foundation models that we have described, GMAI, will interchangeably parse multiple data modalities, learn new tasks on the fly and leverage domain knowledge, offering opportunities across a nearly unlimited range of medical tasks. GMAI’s flexibility allows models to stay relevant in new settings and keep pace with emerging diseases and technologies without needing to be constantly retrained from scratch. GMAI-based applications will be deployed both in traditional clinical settings and on remote devices such as smartphones, and we predict that they will be useful to diverse audiences, enabling both clinician-facing and patient-facing applications.
Despite their promise, GMAI models present unique challenges. Their extreme versatility makes them difficult to comprehensively validate, and their size can bring increased computational costs. There will be particular difficulties associated with data collection and access, as GMAI’s training datasets must be not only large but also diverse, with adequate privacy protections. We implore the AI community and clinical stakeholders to carefully consider these challenges early on, to ensure that GMAI consistently delivers clinical value. Ultimately, GMAI promises unprecedented possibilities for healthcare, supporting clinicians amid a range of essential tasks, overcoming communication barriers, making high-quality care more widely accessible, and reducing the administrative burden on clinicians to allow them to spend more time with patients.
References
- Bommasani, R. et al. On the opportunities and risks of foundation models. Preprint at https://arxiv.org/abs/2108.07258 (2022).
- Reed, S. et al. A generalist agent. In Transactions on Machine Learning Research (2022). This study presented Gato, a generalist model that can carry out a variety of tasks across modalities such as chatting, captioning images, playing video games and controlling a robot arm.
- Alayrac, J.-B. et al. Flamingo: a Visual Language Model for few-shot learning. In Advances in Neural Information Processing Systems (eds Oh, A. H. et al.) 35, 23716–23736 (2022).
- Lu, J., Clark, C., Zellers, R., Mottaghi, R. & Kembhavi, A. Unified-IO: a unified model for vision, language, and multi-modal tasks. Preprint at https://arxiv.org/abs/2206.08916 (2022).
- Brown, T. et al. Language models are few-shot learners. In Advances in Neural Information Processing Systems (eds Larochelle, H. et al.) 33, 1877–1901 (2020). This study presented the language model GPT-3 and discovered that large language models can carry out in-context learning.
- Aghajanyan, A. et al. CM3: a causal masked multimodal model of the Internet. Preprint at https://arxiv.org/abs/2201.07520 (2022).
- Wei, J. et al. Emergent abilities of large language models. In Transactions on Machine Learning Research (2022).
Steinberg, E. et al. Language models are an effective representation learning technique for electronic health record data. J. Biomed. Inform. 113, 103637 (2021).
- Tiu, E. et al. Expert-level detection of pathologies from unannotated chest X-ray images via self-supervised learning. Nat. Biomed. Eng. 6, 1399–1406 (2022). This study demonstrated that CheXzero—an early example of a foundation model in medical AI—can detect diseases on chest X-rays without explicit annotation by learning from natural-language descriptions contained in accompanying clinical reports.
- Singhal, K. et al. Large language models encode clinical knowledge. Preprint at https://arxiv.org/abs/2212.13138 (2022). This study demonstrated that the language model Flan-PaLM achieves a passing score (67.6%) on a dataset of US Medical Licensing Examination questions and proposed Med-PaLM, a medical variant of Flan-PaLM with improved clinical reasoning and comprehension.
Yang, X. et al. A large language model for electronic health records. npj Digit. Med. 5, 194 (2022).
Article PubMed PubMed Central Google Scholar
- Food and Drug Administration. Artificial intelligence and machine learning (AI/ML)-enabled medical devices. FDA https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-and-machine-learning-aiml-enabled-medical-devices (2022).
Acosta, J. N., Falcone, G. J., Rajpurkar, P. & Topol, E. J. Multimodal biomedical AI. Nat. Med. 28, 1773–1784 (2022).
Article CAS PubMed Google Scholar
Krishnan, R., Rajpurkar, P. & Topol, E. J. Self-supervised learning in medicine and healthcare. Nat. Biomed. Eng. 6, 1346–1352 (2022).
- Devlin, J., Chang, M.-W., Lee, K. & Toutanova, K. BERT: pre-training of deep bidirectional transformers for language understanding. In Proc. 2019 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (eds Burstein, J., Doran, C. & Solorio, T.) 1, 4171–4186 (2019). This paper introduced masked language modelling, a widely used technique for training language models where parts of a text sequence are hidden (masked) in order for the model to fill in the blanks. This strategy can be extended beyond text to other data types.
- Radford, A. et al. Learning transferable visual models from natural language supervision. In Proc. 38th Int. Conference on Machine Learning (eds Meila, M. & Zhang, T.) 139, 8748–8763 (2021). This paper introduced contrastive language–image pretraining (CLIP), a multimodal approach that enabled a model to learn from images paired with raw text.
Zhang, X.-A. et al. A zoonotic henipavirus in febrile patients in China. N. Engl. J. Med. 387, 470–472 (2022).
- Vaswani, A. et al. Attention is all you need. In Advances in Neural Information Processing Systems (eds Guyon, I. et al.) 30, 5998–6008 (2017). This paper introduced the transformer architecture, a key breakthrough that ultimately led to the development of large-scale foundation models.
- Borgeaud, S. et al. Improving language models by retrieving from trillions of tokens. In Proc. 39th Int. Conference on Machine Learning (eds Chaudhuri, K. et al.) 162, 2206–2240 (2022).
- Guu, K., Lee, K., Tung, Z., Pasupat, P. & Chang, M.-W. REALM: retrieval-augmented language model pre-training. In Proc. 37th Int. Conference on Machine Learning (eds Daumé, H. & Singh, A.) 119, 3929–3938 (2020).
Igelström, E. et al. Causal inference and effect estimation using observational data. J. Epidemiol. Community Health 76, 960–966 (2022).
Wang, Q., Huang, K., Chandak, P., Zitnik, M. & Gehlenborg, N. Extending the nested model for user-centric XAI: a design study on GNN-based drug repurposing. IEEE Trans. Vis. Comput. Graph. 29, 1266–1276 (2023).
Li, J. et al. Align before fuse: vision and language representation learning with momentum distillation. In Advances in Neural Information Processing Systems (eds Ranzato, M. et al.) 34, 9694–9705 (2021).
- Wang, Z. et al. SimVLM: simple visual language model pretraining with weak supervision. In Int. Conference on Learning Representations (eds Hofmann, K. & Rush, A.) (2022).
- Yasunaga, M. et al. Deep bidirectional language-knowledge graph pretraining. In Advances in Neural Information Processing Systems (eds Oh, A. H. et al.) 35 (2022).
- Yasunaga, M., Ren, H., Bosselut, A., Liang, P. & Leskovec, J. QA-GNN: reasoning with language models and knowledge graphs for question answering. In Proc. 2021 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (eds Toutanova, K. et al.) 535–546 (2021).
Guha Roy, A. et al. Does your dermatology classifier know what it doesn’t know? Detecting the long-tail of unseen conditions. Med. Image Anal. 75, 102274 (2022).
- Radford, A. et al. Robust speech recognition via large-scale weak supervision. Preprint at https://arxiv.org/abs/2212.04356 (2022).
Dixon, R. F. et al. A virtual type 2 diabetes clinic using continuous glucose monitoring and endocrinology visits. J. Diabetes Sci. Technol. 14, 908–911 (2020).
Kucera, T., Togninalli, M. & Meng-Papaxanthos, L. Conditional generative modeling for de novo protein design with hierarchical functions. Bioinformatics 38, 3454–3461 (2022).
Article CAS PubMed PubMed Central Google Scholar
- Rombach, R., Blattmann, A., Lorenz, D., Esser, P. & Ommer, B. High-resolution image synthesis with latent diffusion models. In Proc. IEEE/CVF Conference on Computer Vision and Pattern Recognition (eds Chellappa, R. et al.) 10684–10695 (2022).
- Ramesh, A. et al. Zero-shot text-to-image generation. In Proc. 38th Int. Conference on Machine Learning (eds Meila, M. & Zhang, T.) 139, 8821–8831 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Article ADS CAS PubMed PubMed Central Google Scholar
- Zvyagin, M. et al. GenSLMs: genome-scale language models reveal SARS-CoV-2 evolutionary dynamics. Preprint at bioRxiv https://doi.org/10.1101/2022.10.10.511571 (2022).
- Watson, J. L. et al. Broadly applicable and accurate protein design by integrating structure prediction networks and diffusion generative models. Preprint at bioRxiv https://doi.org/10.1101/2022.12.09.519842 (2022).
The UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 45, D158–D169 (2017).
Guo, L. L. et al. Systematic review of approaches to preserve machine learning performance in the presence of temporal dataset shift in clinical medicine. Appl. Clin. Inform. 12, 808–815 (2021).
Article PubMed PubMed Central Google Scholar
Finlayson, S. G. et al. The clinician and dataset shift in artificial intelligence. N. Engl. J. Med. 385, 283–286 (2021).
Article PubMed PubMed Central Google Scholar
- Lampinen, A. K. et al. Can language models learn from explanations in context? In Findings of the Association for Computational Linguistics: EMNLP 2022 (eds Goldberg, Y., Kozareva, Z. & Zhang, Y.) 537–563 (2022).
Yoon, S. H., Lee, J. H. & Kim, B.-N. Chest CT findings in hospitalized patients with SARS-CoV-2: Delta versus Omicron variants. Radiology 306, 252–260 (2023).
- Ouyang, L. et al. Training language models to follow instructions with human feedback. In Advances in Neural Information Processing Systems (eds Oh, A. H. et al.) 35, 27730–27744 (2022).
- Pilipiszyn, A. GPT-3 powers the next generation of apps. OpenAI https://openai.com/blog/gpt-3-apps/ (2021).
- Burns, C., Ye, H., Klein, D. & Steinhardt, J. Discovering latent knowledge in language models without supervision. Preprint at https://arxiv.org/abs/2212.03827 (2022).
Obermeyer, Z., Powers, B., Vogeli, C. & Mullainathan, S. Dissecting racial bias in an algorithm used to manage the health of populations. Science 366, 447–453 (2019).
Article ADS CAS PubMed Google Scholar
- Sex and Gender Bias in Technology and Artificial Intelligence: Biomedicine and Healthcare Applications (Academic, 2022).
- Srivastava, A. et al. Beyond the imitation game: quantifying and extrapolating the capabilities of language models. Preprint at https://arxiv.org/abs/2206.04615 (2022).
- Carlini, N. et al. Extracting training data from large language models. In Proc. 30th USENIX Security Symposium (eds Bailey, M. & Greenstadt, R.) 6, 2633–2650 (2021).
- Branch, H. J. et al. Evaluating the susceptibility of pre-trained language models via handcrafted adversarial examples. Preprint at https://arxiv.org/abs/2209.02128 (2022).
- Chowdhery, A. et al. PaLM: scaling language modeling with pathways. Preprint at https://arxiv.org/abs/2204.02311 (2022).
- Zhang, S. et al. OPT: open pre-trained transformer language models. Preprint at https://arxiv.org/abs/2205.01068 (2022).
- Hoffmann, J. et al. An empirical analysis of compute-optimal large language model training. In Advances in Neural Information Processing Systems (eds Oh, A. H. et al.) 35, 30016–30030 (2022).
- Chung, H. W. et al. Scaling instruction-finetuned language models. Preprint at https://arxiv.org/abs/2210.11416 (2022).
- Kung, T. H. et al. Performance of ChatGPT on USMLE: potential for AI-assisted medical education using large language models. PLoS Dig. Health 2, 2 (2023).
- Huang, S.-C., Shen, L., Lungren, M. P. & Yeung, S. GLoRIA: a multimodal global-local representation learning framework for label-efficient medical image recognition. In Proc. IEEE/CVF Int. Conference on Computer Vision (eds Brown, M. S. et al.) 3942–3951 (2021).
Johnson, A. E. W. et al. MIMIC-IV, a freely accessible electronic health record dataset. Sci. Data 10, 1 (2023).
Article PubMed PubMed Central Google Scholar
Sudlow, C. et al. UK Biobank: an open access resource for identifying the causes of a wide range of complex diseases of middle and old age. PLoS Med. 12, e1001779 (2015).
Article PubMed PubMed Central Google Scholar
Gou, J., Yu, B., Maybank, S. J. & Tao, D. Knowledge distillation: a survey. Int. J. Comput. Vis. 129, 1789–1819 (2021).
Vegunta, R., Vegunta, R. & Kutti Sridharan, G. Secondary aortoduodenal fistula presenting as gastrointestinal bleeding and fungemia. Cureus 11, e5575 (2019).
PubMed PubMed Central Google Scholar
Acknowledgements
We gratefully acknowledge I. Kohane for providing insightful comments that improved the manuscript. E.J.T. is supported by the National Institutes of Health (NIH) National Center for Advancing Translational Sciences grant UL1TR001114. M.M. is supported by Defense Advanced Research Projects Agency (DARPA) N660011924033 (MCS), NIH National Institute of Neurological Disorders and Stroke R61 NS11865, GSK and Wu Tsai Neurosciences Institute. J.L. was supported by DARPA under Nos. HR00112190039 (TAMI) and N660011924033 (MCS), the Army Research Office under Nos. W911NF-16-1-0342 (MURI) and W911NF-16-1-0171 (DURIP), the National Science Foundation under Nos. OAC-1835598 (CINES), OAC-1934578 (HDR) and CCF-1918940 (Expeditions), the NIH under no. 3U54HG010426-04S1 (HuBMAP), Stanford Data Science Initiative, Wu Tsai Neurosciences Institute, Amazon, Docomo, GSK, Hitachi, Intel, JPMorgan Chase, Juniper Networks, KDDI, NEC and Toshiba.
Author informationAuthor notes
- These authors contributed equally: Michael Moor, Oishi Banerjee
- These authors jointly supervised this work: Eric J. Topol, Pranav Rajpurkar
Authors and Affiliations
Department of Computer Science, Stanford University, Stanford, CA, USA
Michael Moor & Jure Leskovec
Department of Biomedical Informatics, Harvard University, Cambridge, MA, USA
Oishi Banerjee & Pranav Rajpurkar
Institute of Health Policy, Management and Evaluation, Dalla Lana School of Public Health, University of Toronto, Toronto, Ontario, Canada
Zahra Shakeri Hossein Abad
Yale University School of Medicine, Center for Outcomes Research and Evaluation, Yale New Haven Hospital, New Haven, CT, USA
Harlan M. Krumholz
Scripps Research Translational Institute, La Jolla, CA, USA
Eric J. Topol
Contributions
P.R. conceived the study. M.M., O.B., E.J.T. and P.R. designed the review article. M.M. and O.B. made substantial contributions to the synthesis and writing of the article. Z.S.H.A. and M.M. designed and implemented the illustrations. All authors provided critical feedback and substantially contributed to the revision of the manuscript.
Corresponding authors
Correspondence to Eric J. Topol or Pranav Rajpurkar.
Ethics declarations
Competing interests
In the past three years, H.M.K. received expenses and/or personal fees from UnitedHealth, Element Science, Eyedentifeye, and F-Prime; is a co-founder of Refactor Health and HugoHealth; and is associated with contracts, through Yale New Haven Hospital, from the Centers for Medicare & Medicaid Services and through Yale University from the Food and Drug Administration, Johnson & Johnson, Google and Pfizer. The other authors declare no competing interests.
Peer review
Peer review information
Nature thanks Arman Cohan, Joseph Ledsam and Jenna Wiens for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moor, M., Banerjee, O., Abad, Z.S.H. et al. Foundation models for generalist medical artificial intelligence. Nature 616, 259–265 (2023). https://doi.org/10.1038/s41586-023-05881-4